Predictive analysis of metabolic syndrome based on 5-years continuous physical examination data

Diagnostic criteria for MetS
Many researchers around the world have developed clinical criteria for MetS31,32,33,34. To avoid inconsistencies caused by various criteria and best fit the local context of the experimental data, we used the criteria proposed by the Chinese Guidelines for the Prevention and Treatment of Type 2 Diabetes (2017 edition) in this study to identify patients with MetS.
According to the guidelines, patients with MetS can be diagnosed by meeting three or more of the following five conditions.
-
(1)
Abdominal obesity: WC>= 90/85 cm (male/female).
-
(2)
Hyperglycemia: fasting glucose (FGLU)>= 6.1 mmol/L or 2-h postprandial glucose (PG)>= 7.8 mmol/L and/or previously diagnosed and treated diabetes mellitus.
-
(3)
Hypertension: BP>= 130/85 and previously diagnosed and treated hypertension.
-
(4)
Fasting TG>= 1.70 mmol/L.
-
(5)
Fasting HDL-C<= 1.04 mmol/L.
Datasets
We used medical checkup data provided by the Health Management Department of Southern Hospital of Southern Medical University. The dataset contained 1,039,564 medical checkup records for 546,918 participants across 21 prefecture-level cities and subordinate districts and counties in South China, including Guangzhou, Foshan, etc. The inclusion criteria targeted individuals who were 18–80 years old, based on continuous physical examinations taken from 2009 to 2019.
The hospital staff collected numerous raw indicators, including anthropometric data, blood parameters, other biochemical indicators, medical history, gender and age, by extracting values recorded in the physical examination report. Based on the data provided by the hospital, we first derived two additional characteristics from the available anthropometric variables, including the waist-to-hip ratio (WHR) and the body mass index (BMI). Since we aimed to extract new features from specific indicator values that can reflect temporal changes, we extracted 18-dimensional continuous-type numerical features from the data.
After determining the features to be extracted, we cleaned the dataset. We first excluded individuals who underwent too many physical examinations (greater than 20). We then removed outliers for each indicator, including abnormal records for age (greater than 80 years), based on the upper and lower limits of the indicators, as determined by the physicians. After removing the outliers, the treatment of missing values was equally important, as too many missing values can make the model complex. We used different padding strategies for the missing rate, data type and value distribution of each indicator. If the number of missing values was large (more than 70% of data was missing), the feature was deleted. For features with a small number of missing values, we used mean padding if the feature obeyed a normal distribution and median padding if it obeyed a skewed distribution.
After pre-processing the data, we obtained usable structured data containing 530,091 male patients and 398,793 female patients. A more detailed description of these extracted features is shown in Table 6.
This study was approved by the Academic Committee of South China Normal University (Approval No.: SCNU-PHY-2020-063). All methods we used in the study adhered to relevant ethical guidelines and regulations (the Declaration of Helsinki). All patients signed an informed consent form before their data were included in the study.
The MetS prediction model
This study was conducted to compare the differential performance of changes in indicators between those who developed a disease from a healthy state and those who remained healthy. Our findings might help in the effective prevention and intervention of the physically examined population for MetS-related risk factors. We considered the results of five consecutive years of physical examinations for each individual. The features extracted from the multi-year records were used as input to identify the features that could represent physical and physiological changes in the body over time.
The prediction can be considered to be supervised classification, and the first four records used as input in the constructed 5-year model were the features under the healthy state (MS_result = 0). The patients suffering from MetS in the following year were marked as 1 or 0. Therefore, one sample in the model had data on multiple consecutive physical examinations, representing the numerical situation of each index in each year, as shown in Figure 4. After constructing the model, we obtained 15,661 valid samples, of which 1338 and 14,323 samples suffered and did not suffer from MetS in the following year, respectively.
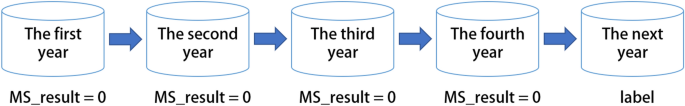
The schematic diagram of the MetS risk prediction model within the next 1 year.
Feature structure
Firstly, we described the year-to-year variation of the indicator values. In this case, the numerical difference feature (DNF) was represented as
$$\beginaligned I\_DNF = I_2 – I_1 \endaligned$$
(1)
where \(I_2\) and \(I_1\) represent the specific values of the indicator in the current year and the past year, respectively. Thus, \(I\_DNF\) could describe the absolute value change of the indicator with the year.
However, it was not enough to have numerical differential features to describe the variation of features in time series. When a patient and an average person changed the exact value of an indicator simultaneously, their importance differed. Therefore, we introduced a weighting function that reflected the different importance when different values were brought to change based on different values.
The weighting function has the following main requirements.
-
(1)
The indicator produces changes in different values, and the risk weight it imposes are different.
-
(2)
The higher the value of each indicator, the higher the risk when generating changes, i.e., the risk weight is an increasing function that grows with the value.
-
(3)
The fastest growth rate of risk is when qualitative changes occur around the upper and lower limits of the indicator.
We obtained by observing different function algorithms that the underlying formula of the Sigmoid function meets our requirements. The image of the Sigmoid function is continuous and smooth, strictly monotonic, and symmetric with a (0,0.5) center. Therefore, the risk growth curve could be described as
$$\beginaligned S(x) = 1 / (1 + e^-ax) \endaligned$$
(2)
where x is the difference between the current indicator value and the upper limit of the normal range of that indicator, and a is a function parameter.
According to the above formulas, the new feature was defined as the product of the numerical change weight function and the numerical difference feature of the indicator, as shown in
$$\beginaligned F(x) = I\_DNF * S(x) \endaligned$$
(3)
where the function S(x) is a weight function modeled after the Sigmoid function, and \(I\_DNF\) is the difference between the values for a certain 2-year period. The result is expressed as the effect of the disease brought about by a specific 2-year change at different values.
Since the study was conducted on a sample set of consecutive years, we obtained three new features for each indicator, i.e., new features generated by the above formula for years 1-2, 2-3, and 3-4. To effectively combine the three time periods, we gave different weights to the new features for each period, where the new features closer to the current time have higher weights, which were determined using the formula \(b^3 +b^2 +b^1=1\). Finally, the formula of new features was shown as
$$\beginaligned G(x_1,x_2,x_3)=b^3*I_1\_DNF*S(x_1)+b^2*I_2\_DNF*S(x_2)+b*I_3\_DNF*S(x_3) \endaligned$$
(4)
where \(x_1\), \(x_2\) and \(x_3\) are the differences between the values and the normal limit in the past every year. Moreover, \(I_1\_DNF\), \(I_2\_DNF\) and \(I_3\_DNF\) are the differences between the values in years 1-2, 2-3, and 3-4.
We reconciled parameters a and b in subsequent work based on cross-validation of machine learning to achieve optimal performance. Since each indicator had taken different value ranges, parameter a’s significance was to scale the value ranges of different indicators uniformly. Therefore, we normalized the maximum value of x by (1/maximum value of different indicators) and then gave the parameter value \(a=50\) which performed best in machine learning according to different scaling degrees. For parameter b, the optimal parameter \(b=0.6\) was selected based on the exact weight formula as above with the performance of machine learning.
Machine learning classifier
We used several algorithms to observe the performance of predictive models in machine learning for 5 years of data, including the more commonly used machine learning algorithms and the traditional logistic regression algorithm.
-
(1)
XGBoost: XGBoost consists of several decision trees, of which the decision tree is a CART regression tree model. The main idea of this classifier is to continuously learn new regression trees to fit the residuals of the last prediction, thus obtaining very high accuracy.
-
(2)
Random Forest (RF): RF is a common machine learning classifier based on the bagging algorithm35, which also consists of a combination of multiple decision trees. Compared with the traditional single-tree classifier, RF has a fairly high-performance optimization.
-
(3)
Stacking: Stacking is an integrated classifier based on adding another layer of classifiers on top of the original classifier, then selecting the target labels predicted by most of the classifiers by a voting method. In this paper, we let Stacking combine XGBoost and RF.
-
(4)
Logistic Regression (LR): LR is a traditional classical classifier, which works similar to linear regression, assuming that the data obeys a certain distribution and then using a great likelihood estimation algorithm to do parameter estimation.
Evaluation
In our experiments, the primary metric used to determine the classifier’s performance was the Area Under The Curve (AUC) of the subject’s operating characteristic curve (ROC). The AUC with superior discriminative ability is 1.0, and the AUC without discriminative ability is 0.5. To comprehensively consider the performance of machine learning classifiers, we also used the metrics Accuracy, Sensitivity, Specificity, Precision and F1-score for evaluation, defined as
$$\beginaligned Accuracy= & (TP+TN)/(TP+FP+FN+TN) \endaligned$$
(5)
$$\beginaligned Precision= & TP/(TP+FP) \endaligned$$
(6)
$$\beginaligned Sensitivity= & TP/(TP+FN) \endaligned$$
(7)
$$\beginaligned Specificity= & TN/(TN+FP) \endaligned$$
(8)
$$\beginaligned F1\_score= & 2*(Precision*Sensitivity)/(Precision+Sensitivity) \endaligned$$
(9)
where TP (true positive), TN (true negative), FP (false positive) and FN (false negative) are the values in the confusion matrix, and each final result was subjected to multiplicative cross-validation. Then we took their mean and standard deviation.
link



![Average Cost of Medical School [2025]: Yearly + Total Costs Average Cost of Medical School [2025]: Yearly + Total Costs](https://educationdata.org/wp-content/uploads/2025/09/page-1.png)