Multi-layer encrypted learning for distributed healthcare analytics
This paper proposes a novel privacy-preserving framework designed with a three-layer protection mechanism for distributed healthcare analytics on encrypted data. The proposed framework comprises the following three components:
Computation on encrypted data
FHE enables analytical computations to be performed directly on encrypted data. Thus, the proposed framework mitigates the risk of data breaches and preserves patient privacy throughout the analytical process.
Decentralized on‑site learning
The proposed framework employs decentralized model training across multiple healthcare units without the need to exchange raw data. Each unit trains its model locally and shares only model updates with a central server. This federated learning design in smart healthcare systems ensures that sensitive patient data remain confidential while still enabling collaborative analytics.
Collective intelligence of models
To enhance computational efficiency and model performance, ensemble learning is utilized within a parallel computing architecture. Multiple sub-models are trained simultaneously and then integrated into a consensus model. This parallelism accelerates the computational process and provides scalable analysis of large-scale healthcare data.
As shown in Fig. 7, the interactive workflow of these layers ensures that raw data are never exchanged between units, while still allowing continuous model refinement and enhanced predictive performance. Integrated interaction between data segregation, FHE encryption, and federated ensemble learning forms the backbone of a privacy-preserving framework for healthcare analytics. The workflow is determined by problem constraints in distributed healthcare system.
Segregated data storage
In the context of IoMT, healthcare data are collected from diverse sources, including wearable devices, sensors, and medical equipment, and are distributed across different units or departments of healthcare systems. Notably, a single unit only provides care to a small number of patients, while different units often treat diverse patient populations. For instance, one pediatric unit may primarily handle a small pool of cases, while the other focuses on specific patients with chronic conditions. This variation results in distributed data that reflect only specific patient populations. As such, units are left with a limited perspective on broader healthcare trends and patient conditions. To obtain a comprehensive view of healthcare trends and patient conditions, collaboration among multiple units across healthcare systems is needed. However, such collaboration introduces significant concerns regarding data privacy, especially when sharing sensitive patient data.
Data storage is crucial for protecting data privacy and enabling efficient operations. Currently, segregated data storage is a common structure where each unit or department across healthcare systems retains full control over its dataset. Data are stored independently. This structure supports a high level of data privacy protection because each unit enforces its privacy protocols to minimize the risk of unauthorized access. However, while segregated data storage helps protect data privacy, the potential for data-driven collaboration across multiple independent healthcare systems is limited. Units face challenges in coordinating care because isolated datasets prevent comprehensive patient records from being assembled. Such limitations reduce insights that could improve diagnostics, treatments, and patient outcomes. As a result, while effective for privacy, segregated data storage hinders the broader goals of an integrated healthcare system.
On the other hand, aggregated data storage is another structure where multiple units across healthcare systems contribute to a unified, collaboratively managed dataset. In this setup, data storage responsibilities are shared among all participating units or departments. This structure enables access to a more extensive dataset that can reveal patterns and insights across patient populations. Aggregated data storage supports more effective disease tracking, personalized treatment strategies, and improved diagnostics. However, this structure poses unique challenges in maintaining patient privacy because of the increasing level of complexity when multiple units have access to the same dataset. Without privacy-preserving mechanisms such as data encryption, anonymization techniques, and controlled access protocols, patients’ sensitive data are at risk. The balance between maximizing the value of shared data and protecting data privacy remains a pressing concern in IoMT.
To address these challenges, it is imperative to develop a privacy-preserving framework for distributed learning in healthcare systems. Such a framework can enable units across healthcare systems to collaborate on data analytics while protecting sensitive patient data. As shown in Fig. 8, the proposed privacy-preserving framework consists of three-layer protection. Patient data from various devices and sensors are standardized and de-identified at each healthcare unit to ensure that data remain localized and compliant with privacy requirements. (Layer 1: Data Segregation). These segregated data are then encrypted by FHE to enable analytical computation without decryption and mitigate the likelihood of data breaches (Layer 2: Data Encryption). The encryption layer interacts with the decentralized learning layer, where federated learning is employed to collaboratively train machine learning models across multiple units. In this process, each unit processes its encrypted data locally and transmits only model updates to a central server. These updates are then integrated using ensemble learning techniques, which combine the strengths of multiple sub-models into a consensus model. (Layer 3: Decentralized Learning). The interactive workflow of these layers ensures that raw data are never exchanged between units, while still allowing continuous model refinement and enhanced predictive performance. Integrated interaction between data segregation, FHE encryption, and federated ensemble learning forms the backbone of a privacy-preserving framework for healthcare analytics.
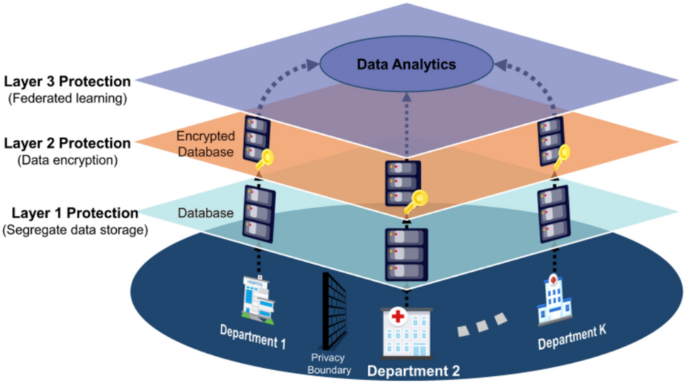
Three-layer privacy protection of the proposed framework.
Computation on encrypted data
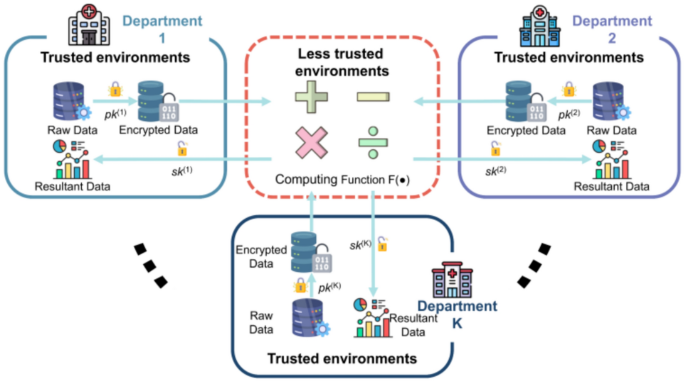
FHE, a novel encryption technique, goes a step further by allowing computations to be conducted directly on encrypted data. As a form of asymmetric encryption, two distinct keys are utilized: a public key (\(pk\)) and a private key (\(sk\)). Notably, the public key is to encrypt the raw data, and the private key is to decrypt it. A significant advantage of FHE is that, upon decryption, the results of these computations are identical to those obtained by performing on raw data. This capability allows data processing and analysis while preserving data privacy.
As shown in Fig. 9, there are K departments in the healthcare system. Department \(k\) independently manages its data \({\left[\mathbf{X},\mathbf{Y}\right]}^{\left(k\right)}\), where \(\mathbf{X}\) denotes the matrix of input variables, and \(\mathbf{Y}\) is the matrix of output values in the process of healthcare analytics. Each department \(k\) owns its key pair, including the public key \({pk}^{\left(k\right)}\) and the private key \({sk}^{\left(k\right)}\). Upon data collection, department \(k\) encrypts the raw data using \({pk}^{\left(k\right)}\) into \({\left[{\mathbf{X}}_{e} , {\mathbf{Y}}_{e}\right]}^{\left(k\right)}\). Notably, this encrypted data can only be decrypted by department \(k\), which holds the corresponding private key \({sk}^{\left(k\right)}\). When department \(k\) conducts analytical computation such as addition or multiplication on encrypted data, FHE ensures that the operations yield accurate results. For instance, if an addition operation is defined as \(F\left({x}_{e},{ y}_{e}\right)\text{=}{ x}_{e}\text{+}{y}_{e}\), decrypting the result with the \({sk}^{\left(k\right)}\) produces the same outcome as \(x\text{+}y\). Similarly, for a multiplication operation defined as \(F\left( {x_{e} , y_{e} } \right){ = } x_{e} \times y_{e}\), decrypting the result yields an outcome identical to \(x \times y\).
In the proposed framework, each unit or department \(k\) of healthcare systems generates a key pair, including a private key \({sk}^{\left(k\right)}\) and a public key \({pk}^{\left(k\right)}\), following the assumption of Ring Learning with error (RLWE)28. Define
$$\begin{array}{*{20}c} {R{ = }{\raise0.7ex\hbox{${{\mathbb{Z}}\left[ X \right]}$} \!\mathord{\left/ {\vphantom {{{\mathbb{Z}}\left[ X \right]} {\left( {X^{n} { + }1} \right)}}}\right.\kern-0pt} \!\lower0.7ex\hbox{${\left( {X^{n} { + }1} \right)}$}}} \\ \end{array}$$
(1)
as the cyclotomic ring, where \(n\) is a power of two and \({\mathbb{Z}}\left[X\right]\) is defined as the polynomial ring with integer coefficients. The residue ring \({R}_{q}\text{=}{\mathbb{Z}}_{q}\left[X\right]/\left({X}^{n}\text{+}1\right)\) operates with coefficients modulo \(q\). Key generation utilizes polynomials in the form \(\left( {a^{\left( k \right)} ,b^{\left( k \right)} { = – }s^{\left( k \right)} \cdot a^{\left( k \right)} { + }e} \right)\) . Per the RLWE assumption, \({b}^{\left(k\right)}\) is indistinguishable from uniformly random elements in \({R}_{q}\) when \({a}^{\left(k\right)}\) is selected uniformly at random from \({R}_{q}\), \({s}^{\left(k\right)}\) is drawn from \({R}_{q}\), and \(e\) is sampled from a uniform distribution over \({R}_{q}\). The fundamental settings of FHE algorithm are outlined below.
-
Private key setup: For each unit \(k\), the private key \({sk}^{\left(k\right)}\) is defined as:
$$\begin{array}{c}{sk}^{\left(k\right)}\text{=}\left(1,{s}^{\left(k\right)}\right)\#\end{array}$$
(2)
-
Public key setup: The public key \({pk}^{\left(k\right)}\) of unit \(k\) is defined by the following equation:
$$\begin{array}{*{20}c} {pk^{\left( k \right)} { = }\left( {b^{\left( k \right)} ,a^{\left( k \right)} } \right){ = }\left( {{ – }s^{\left( k \right)} \cdot a^{\left( k \right)} { + }e,a^{\left( k \right)} } \right)} \\ \end{array}$$
(3)
-
Encryption \({\text{Enc}}\left(\cdot \right)\): The FHE takes a raw data \({x}^{\left(k\right)}\) and the public key \({pk}^{\left(k\right)}\) from unit \(k\) as input and outputs an encrypted data \({x}_{e}^{\left(k\right)}\) as follows:
$$\begin{aligned} x_{e}^{\left( k \right)} { = } & \,{\text{Enc}}\left( {pk^{\left( k \right)} ,x^{\left( k \right)} } \right) = \left( {x^{\left( k \right)} ,0} \right){ + }pk^{\left( k \right)} \\ = & \left( {x^{\left( k \right)} { – }s^{\left( k \right)} \cdot a^{\left( k \right)} { + }e,a^{\left( k \right)} } \right) \\ = & \left( {c_{0}^{\left( k \right)} ,c_{1}^{\left( k \right)} } \right) \\ \end{aligned}$$
(4)
-
Decryption \({\text{Dec}}\left(\cdot \right)\): The FHE decryption process takes the encrypted data \({x}_{e}^{\left(k\right)}\) and the private key \({sk}^{\left(k\right)}\) as input and generates a decrypted data \({\widetilde{x}}^{\left(k\right)}\) as follow:
$$\begin{aligned} \tilde{x}^{\left( k \right)} = &\, {\text{Dec}}\,\left( {sk^{\left( k \right)} ,x_{e}^{\left( k \right)} } \right) \\ = &\, c_{0}^{\left( k \right)} { + }c_{1}^{\left( k \right)} \cdot s^{\left( k \right)} \\ = & \left( {x^{\left( k \right)} { – }s^{\left( k \right)} \cdot a^{\left( k \right)} { + }e} \right){ + }a^{\left( k \right)} \cdot s^{\left( k \right)} \\ = & x^{\left( k \right)} { + }e \\ \approx & x^{\left( k \right)} \\ \end{aligned}$$
(5)
Two key properties of FHE are described below:
-
Additive homomorphism \({\text{Add}}\left(\cdot \right)\): Suppose unit \(k\) has two data point \({x}^{\left(k\right)}\) and \({y}^{\left(k\right)}\), which are encrypted as follows:
$$\begin{array}{c}{x}_{e}^{\left(k\right)}\text{=}\,{\text{E}}{\text{n}}{\text{c}}\left({pk}^{\left(k\right)},{x}^{\left(k\right)}\right)\text{=}\left({c}_{x,0}^{\left(k\right)},{c}_{x,1}^{\left(k\right)}\right)\end{array}$$
(6)
$$\begin{array}{c}{y}_{e}^{\left(k\right)}\text{=}{\text{E}}{\text{n}}{\text{c}}\left({pk}^{\left(k\right)},{y}^{\left(k\right)}\right)\text{=}\left({c}_{y,0}^{\left(k\right)},{c}_{y,1}^{\left(k\right)}\right)\end{array}$$
(7)
to mitigate the risk of data breaches while computing data analysis. When an addition operation is demanded, the encrypted sum of \({x}_{e}^{\left(k\right)}\) and \({y}_{e}^{\left(k\right)}\) is calculated as follows:
$$\begin{aligned} {\text{Add}}\left( {x_{e}^{\left( k \right)} ,y_{e}^{\left( k \right)} } \right) = & x_{e}^{\left( k \right)} { + }y_{e}^{\left( k \right)} \\ = & \left( {c_{x,0}^{\left( k \right)} ,c_{x,1}^{\left( k \right)} } \right){ + }\left( {c_{y,0}^{\left( k \right)} ,c_{y,1}^{\left( k \right)} } \right) \\ = & \left( {c_{x,0}^{\left( k \right)} { + }c_{y,0}^{\left( k \right)} ,c_{x,1}^{\left( k \right)} { + }c_{y,1}^{\left( k \right)} } \right) \\ \end{aligned}$$
(8)
When the result of \({\text{Add}}\left({x}_{e}^{\left(k\right)}\text{+}{y}_{e}^{\left(k\right)}\right)\) is decrypted by \({sk}^{\left(k\right)}\), the outcome is the same as the addition of raw data \({x}^{\left(k\right)}\text{+}{y}^{\left(k\right)}\), as shown below:
$$\begin{aligned} {\text{Dec}} & \left( {sk^{\left( k \right)} ,{\text{Add}}\left( {x_{e}^{\left( k \right)} ,y_{e}^{\left( k \right)} } \right)} \right) = \left( {c_{x,0}^{\left( k \right)} + c_{y,0}^{\left( k \right)} } \right){ + }\left( {c_{x,1}^{\left( k \right)} + c_{y,1}^{\left( k \right)} } \right) \cdot sk^{\left( k \right)} \\ = & \left( {c_{x,0}^{\left( k \right)} + c_{x,1}^{\left( k \right)} \cdot sk^{\left( k \right)} } \right) + \left( {c_{y,0}^{\left( k \right)} + c_{y,1}^{\left( k \right)} \cdot sk^{\left( k \right)} } \right) \\ = & \left( {x^{\left( k \right)} + e} \right) + \left( {y^{\left( k \right)} + e} \right) \approx x^{\left( k \right)} + y^{\left( k \right)} \\ \end{aligned}$$
(9)
-
Multiplicative homomorphism \({\text{Multi}}\left(\cdot \right)\): Conversely, if unit \(k\) has one encrypted data \({x}_{e}^{\left(k\right)}\) and performs multiplication by a non-sensitive real number \(t\), which does not need encryption, the multiplication function is defined as follows:
$$\begin{aligned} {\text{Multi}}\left( {x_{e}^{\left( k \right)} ,t} \right) =\, & x_{e}^{\left( k \right)} \cdot t \\ = & \left( {c_{x,0}^{\left( k \right)} ,c_{x,1}^{\left( k \right)} } \right) \cdot t \\ = & \left( {c_{x,0}^{\left( k \right)} \cdot t,c_{x,1}^{\left( k \right)} \cdot t} \right) \\ \end{aligned}$$
(10)
When the private key \({sk}^{\left(k\right)}\) is utilized to decrypt this result, the decryption yields:
$$\begin{aligned} {\text{Dec}}\left( {sk^{\left( k \right)} ,{\text{Multi}}\left( {x_{e}^{\left( k \right)} ,t} \right)} \right) =\, & c_{x,0}^{\left( k \right)} \cdot t{ + }c_{x,1}^{\left( k \right)} \cdot t \cdot sk^{\left( k \right)} \\ =\, & t \cdot \left( {c_{x,0}^{\left( k \right)} { + }c_{x,1}^{\left( k \right)} \cdot sk^{\left( k \right)} } \right) \\ =\, & t \cdot \left( {x^{\left( k \right)} { + }e} \right) \\ =\, & t \cdot x^{\left( k \right)} { + }t \cdot e \\ \approx\, & t \cdot x^{\left( k \right)} \\ \end{aligned}$$
(11)
Furthermore, if data \({x}^{\left(k\right)}\) and \({y}^{\left(k\right)}\) are sensitive and used in the multiplication function, unit \(k\) has to encrypt them as \({x}_{e}^{\left(k\right)}\) and \({y}_{e}^{\left(k\right)}\). The multiplication function for two encrypted data is defined as follows:
$$\begin{aligned} {\text{Dec}} & \left( {sk^{\left( k \right)} ,{\text{Multi}}\left( {x_{e}^{\left( k \right)} ,y_{e}^{\left( k \right)} } \right)} \right) \\ = &\, {\text{Dec}}\left( {sk^{\left( k \right)} ,x_{e}^{\left( k \right)} } \right) \cdot {\text{Dec}}\left( {sk^{\left( k \right)} ,y_{e}^{\left( k \right)} } \right) \\ = & \left( {c_{x,0}^{\left( k \right)} { + }c_{x,1}^{\left( k \right)} \cdot s^{\left( k \right)} } \right) \cdot \left( {c_{y,0}^{\left( k \right)} { + }c_{y,1}^{\left( k \right)} \cdot s^{\left( k \right)} } \right) \\ = &\, c_{x,0}^{\left( k \right)} \cdot c_{y,0}^{\left( k \right)} { + }\left( {c_{x,0}^{\left( k \right)} \cdot c_{y,1}^{\left( k \right)} { + }c_{x,1}^{\left( k \right)} \cdot c_{y,0}^{\left( k \right)} } \right) \cdot s^{\left( k \right)} { + }c_{x,1}^{\left( k \right)} \cdot c_{y,1}^{\left( k \right)} \cdot s^{\left( k \right)2} \\ = &\, d_{0} { + }d_{1} \cdot s^{\left( k \right)} { + }d_{2} \cdot s^{\left( k \right)2} \\ \end{aligned}$$
(12)
Here, it is important to note that \({\text{Dec}}\left(\cdot \right)\) is typically a quadratic polynomial. However, as shown in the above equation, \({\text{Dec}}\left({sk}^{\left(k\right)},{\text{Multi}}\left({x}_{e}^{\left(k\right)},{y}_{e}^{\left(k\right)}\right)\right)\) now becomes a cubic polynomial. As a result, relinearization \({\text{ReLin}}\left(\cdot \right)\)29 is adopted to make \({\text{Dec}}\left({sk}^{\left(k\right)},{\text{Multi}}\left({x}_{e}^{\left(k\right)},{y}_{e}^{\left(k\right)}\right)\right)\) still be a quadratic polynomial. The relinearization process is defined as follows:
$$\begin{array}{c}\left({d}_{0}^{\prime},{d}_{1}^{\prime}\right)={\text{R}}{\text{e}}{\text{L}}{\text{i}}{\text{n}}\left({\text{Multi}}\left({x}_{e}^{\left(k\right)},{y}_{e}^{\left(k\right)}\right)\right)\end{array}$$
(13)
where \(d_{0}^{\prime } { + }d_{1}^{\prime } \cdot s^{\left( k \right)} { = }d_{0} { + }d_{1} \cdot s^{\left( k \right)} { + }d_{2} \cdot s^{\left( k \right)2}\). Consequently, the decryption of relinearized result yields:
$$\begin{array}{c}{\text{D}}{\text{e}}{\text{c}}\left({sk}^{\left(k\right)},{\text{ReLin}}\left({\text{Multi}}\left({x}_{e}^{\left(k\right)},{y}_{e}^{\left(k\right)}\right)\right)\right)\approx {x}^{\left(k\right)} {y}^{\left(k\right)}\end{array}$$
(14)
Federated learning and predictive analytics
IoMT leverages vast, diverse, and high-quality data to enhance healthcare capabilities. However, collaboration among multiple units or departments introduces a risk of data breaches. To mitigate this risk, a federated learning framework offers an effective solution by circumventing the need for centralized data storage. Models are trained locally at each unit to keep data securely within its original environment. This decentralized framework enhances data privacy while still supporting the benefits of collaborative learning. This research introduces a federated learning framework tailored to the IoMT context to increase data utility while protecting privacy.
As shown in Fig. 10, a network of \(\text{K}\) departments collaborate across healthcare systems. Each department stores its own dataset, denoted as \({\mathcal{D}}^{\left(k\right)}\text{=}\left({\mathbf{x}}_{i}^{\left(k\right)},{y}_{i}^{\left(k\right)}\right)\), where \(i\text{=}1,\dots ,{N}^{\left(k\right)}\). Here, \({\mathbf{x}}_{i}^{\left(k\right)}\) represents the \({i}^{th}\) input vector for department \(k\), \({y}_{i}^{\left(k\right)}\) corresponds to the \({i}^{th}\) output, and \({N}^{\left(k\right)}\) specifies the total amount of data at department \(k\). Each department securely stores its data and further develops machine learning models. Variations in patient demographics, hospital resources, and other factors contribute to differences in datasets across departments. Subsequently, a consensus model synthesizes the insights from these individual models to create a unified machine learning model.
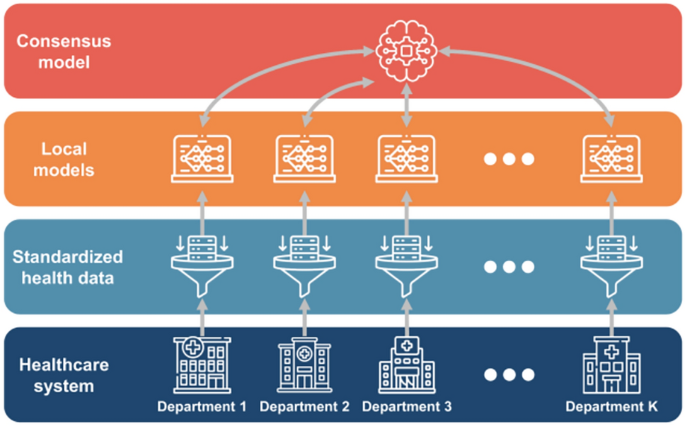
The framework of federated learning in IoMT.
In a multi-entity collaboration setting, this research illustrates the federated learning framework by utilizing logistic regression as an example in healthcare systems. First, each unit or department encrypts its raw data as follows:
$$\begin{array}{c}{\text{E}}{\text{n}}{\text{c}}\left({pk}^{\left(k\right)},{\mathbf{x}}^{\left(k\right)}\right)\text{=}{\mathbf{x}}_{e}^{\left(k\right)}, {\text{E}}{\text{n}}{\text{c}}\left({pk}^{\left(k\right)},{y}^{\left(k\right)}\right)\text{=}{y}_{e}^{\left(k\right)}\end{array}$$
(15)
Here, the logistic regression is defined as
$$\begin{array}{c}log\left(\frac{p}{1-p}\right)\text{=}{\mathbf{x}}_{e}^{\left(k\right)}{{{\varvec{\upbeta}}}^{\left(t-1\right)}}^{\text{T}}\end{array}$$
(16)
where \(p\) represents the probability of outcome \(y\text{=}1\), \({\mathbf{x}}_{e}^{\left(k\right)}\) is the encrypted input vector from the \({k}^{th}\) unit, \({{\varvec{\upbeta}}}^{\left(t-1\right)}\) denotes the parameters vector updated for \(t-1\) times. Notably, \(p\left({\mathbf{x}}_{e}^{\left(k\right)},{\varvec{\upbeta}}\right)\) is defined as the probability of outcome \(y\text{=}1\) on the given encrypted input \({\mathbf{x}}_{e}^{\left(k\right)}\) and the model parameters \({\varvec{\upbeta}}\). After rearranging the above equation, we can have
$$\begin{array}{c}\frac{p}{1-p}\,\,\text{=}\,\,{e}^{{\mathbf{x}}_{e}^{\left(k\right)}{{{\varvec{\upbeta}}}^{\left(t-1\right)}}^{\text{T}}}\end{array}$$
(17)
$$\begin{array}{c}p\,\,\text{=}\,\,\frac{{e}^{{\mathbf{x}}_{e}^{\left(k\right)}{{{\varvec{\upbeta}}}^{\left(t-1\right)}}^{\text{T}}}}{1\text{+}{e}^{{\mathbf{x}}_{e}^{\left(k\right)}{{{\varvec{\upbeta}}}^{\left(t-1\right)}}^{\text{T}}}}\end{array}$$
(18)
$$\begin{array}{c}1-p\,\,\text{=}\,\,\frac{1}{1\text{+}{e}^{{\mathbf{x}}_{e}^{\left(k\right)}{{{\varvec{\upbeta}}}^{\left(t-1\right)}}^{\text{T}}}}\end{array}$$
(19)
The model parameters are estimated through maximum likelihood estimation (MLE), defined as the joint probability density of healthcare data from unit \(k\) conditioned on a given set of model parameters. Therefore, the joint likelihood function for the training data from unit \(k\) is defined as
$$\begin{aligned} {\text{likelihood}}\left( {\varvec{\upbeta}} \right) = & \mathop \prod \limits_{{n:y_{n} = 1}} { }p\left( {{\mathbf{x}}_{e,n}^{\left( k \right)} ,{\varvec{\upbeta}}} \right)\mathop \prod \limits_{{n:y_{n} = 0}} { }\left[ {1{ – }p\left( {{\mathbf{x}}_{e,n}^{\left( k \right)} ,{\varvec{\upbeta}}} \right)} \right] \\ = & \mathop \prod \limits_{n = 1}^{{N^{\left( k \right)} }} p\left( {{\mathbf{x}}_{e,n}^{\left( k \right)} ,{\varvec{\upbeta}}} \right)^{{y_{n} }} \left[ {1{ – }p\left( {{\mathbf{x}}_{e,n}^{\left( k \right)} ,{\varvec{\upbeta}}} \right)} \right]^{{1{ – }y_{n} }} \\ \end{aligned}$$
(20)
Taking the logarithm transforms products into sums as follows:
$$\begin{aligned} l\left( \upbeta \right) = & \log likelihood\left( \upbeta \right) \\ = & \mathop \sum \limits_{{n = 1}}^{{N^{{\left( k \right)}} }} \left[ {y_{n} \log p\left( {x_{{e,n}}^{{\left( k \right)}} ,\upbeta } \right) + \left( {1 – y_{n} } \right)\log \left( {1 – p\left( {x_{{e,n}}^{{\left( k \right)}} ,\upbeta } \right)} \right)} \right] \\ = & \mathop \sum \limits_{{n = 1}}^{{N^{{\left( k \right)}} }} y_{n} x_{{e,n}}^{{\left( k \right)}} \upbeta ^{T} – \log \left( {1 + e^{{x_{{e,n}}^{{\left( k \right)}} \upbeta ^{T} }} } \right)~ \\ \end{aligned}$$
(21)
The objective of MLE is to determine the optimal parameter set \({{\varvec{\upbeta}}}^{\boldsymbol{*}}\) of model that maximizes the log-likelihood function:
$$\begin{array}{c}\underset{{{\varvec{\upbeta}}}^{\boldsymbol{*}}}{\text{max}}l\left({\varvec{\upbeta}}\right)=\underset{{{\varvec{\upbeta}}}^{\boldsymbol{*}}}{\text{max}} \sum_{n=1}^{{N}^{\left(k\right)}}{y}_{n}{\mathbf{x}}_{e,n}^{\left(k\right)}{{\varvec{\upbeta}}}^{\text{T}}-\text{log}\left(1\text{+}{e}^{{\mathbf{x}}_{e,n}^{\left(k\right)}{{\varvec{\upbeta}}}^{\text{T}}}\right)\end{array}$$
(22)
Model parameters \({\varvec{\upbeta}}\) are iteratively updated as follows:
$$\begin{gathered} \upbeta_{0}^{t} = \upbeta_{0}^{t – 1} – \alpha \frac{\partial \upbeta }{{\partial \upbeta_{0} }} = \upbeta_{0}^{t – 1} – \alpha \frac{1}{{N^{\left( k \right)} }}\mathop \sum \limits_{n = 1}^{{N^{\left( k \right)} }} p\left( {x_{e,n}^{\left( k \right)} ,\upbeta } \right) – y_{n} \hfill \\ \upbeta_{1}^{t} = \upbeta_{1}^{t – 1} – \alpha \frac{\partial \upbeta }{{\partial \upbeta_{1} }} = \upbeta_{1}^{t – 1} – \alpha \frac{1}{{N^{\left( k \right)} }}\mathop \sum \limits_{n = 1}^{{N^{\left( k \right)} }} \left( {p\left( {x_{e,n}^{\left( k \right)} ,\upbeta } \right) – y_{n} } \right) \cdot x_{e,n,1} \hfill \\ \upbeta_{W}^{t} = \upbeta_{W}^{t – 1} – \alpha \frac{\partial \upbeta }{{\partial \upbeta_{W} }} = \upbeta_{W}^{t – 1} – \alpha \frac{1}{{N^{\left( k \right)} }}\mathop \sum \limits_{n = 1}^{{N^{\left( k \right)} }} \left( {p\left( {x_{e,n}^{\left( k \right)} ,\upbeta } \right) – y_{n} } \right) \cdot x_{e,n,W} \hfill \\ \end{gathered}$$
(23)
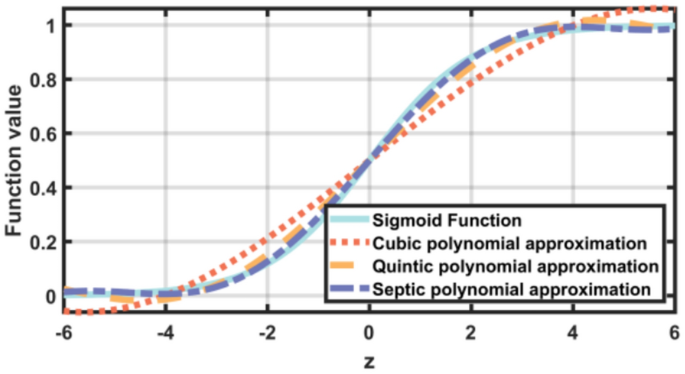
Due to the constraints of FHE, directly implementing the Sigmoid function is infeasible. To address this challenge, a polynomial approximation \({\sigma }_{r}\left(z\right)\) is employed to approximate the sigmoid function:
$$\sigma_{r} \left( z \right) \approx \frac{1}{{1{ + }e^{ – z} }}$$
(24)
where \(r\) represents the polynomial’s order, and \(z\) is the input to the Sigmoid function. The polynomial approximation allows the use of the Sigmoid function within FHE constraints. As such, the proposed framework can support secure computation while preserving the essential properties of the original function.
Ensemble learning in smart healthcare systems
FHE provides a high level of data privacy protection by enabling computations directly on encrypted data. However, this privacy protection comes with a significant computational burden due to the intensive mathematical operations involved. To address this trade-off, a distributed ensemble learning structure is designed in the proposed framework to both reduce computation time and enhance model performance.
Ensemble learning combines multiple sub-models to create a more robust machine learning model. Specifically, bootstrap aggregating (bagging) is employed to enhance model performance by integrating outcomes from several sub-models. In bagging, multiple versions of models are trained on different bootstrap samples of training data, and their outcomes are integrated to produce a final result. This distributed learning structure reduces the computational load for each sub-model by distributing the training process across multiple nodes. By leveraging distributed computing resources, the overall computation time can be reduced, as each model is trained in parallel. Additionally, using various training sets improves model accuracy and robustness by reducing variance and minimizing overfitting.
As shown in Fig. 11, we propose a distributed ensemble learning framework for smart healthcare systems. In this framework, \(\text{K}\) departments collaborate, and the consensus model comprises \(\text{S}\) sub-models. Each department’s dataset is divided into \(\text{S}\) subsets, corresponding to the number of models in the consensus framework. Each subset of data trains a specific sub-model; for instance, subset \(s\) from department \(k\) is used to train sub-model \(s\). As a result, this approach allows each sub-model to process only \(1/s\) of the total data, distributing the computational load across multiple nodes. Once all sub-models are trained, their outputs are combined using a majority voting mechanism to determine the final prediction.
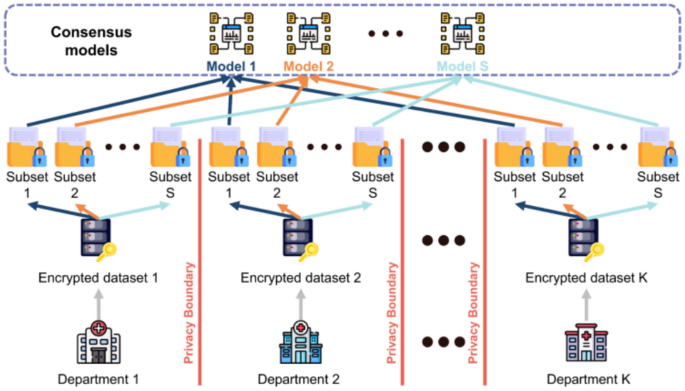
The framework of ensemble learning within the cooperation across multiple departments.
Each logistic regression sub-model computes the probability of the positive class using the sigmoid function:
$$p_{s} \left( {{\mathbf{x}}_{{\varvec{e}}} } \right) = \frac{{e^{{{\mathbf{x}}_{{\varvec{e}}} {\varvec{\upbeta}}_{{\varvec{s}}}^{{\text{T}}} }} }}{{1{ + }e^{{{\mathbf{x}}_{{\varvec{e}}} {\varvec{\upbeta}}_{s}^{{\text{T}}} }} }}$$
(25)
where \({{\varvec{\upbeta}}}_{s}\) denotes the parameters vector for sub-model \(s\). Each sub-model assigns a binary label based on a threshold, which is typically defined as 0.5:
$$\begin{array}{*{20}c} {\hat{y}_{s} = \left\{ {\begin{array}{*{20}c} {1,} & {{\text{if}}\, p_{s} \left( {{\mathbf{x}}_{{\varvec{e}}} } \right) \ge 0.5 } \\ {0,} & {{\text{if}}\, p_{s} \left( {{\mathbf{x}}_{{\varvec{e}}} } \right) < 0.5 } \\ \end{array} } \right. } \\ \end{array}$$
(26)
The final ensemble prediction \({\widehat{y}}_{ens}\) is determined via majority voting as
$$\hat{y}_{ens} = \left\{ {\begin{array}{*{20}c} {1,} & {{\text{if}} \mathop \sum \limits_{s = 1}^{{\text{S}}} \hat{y}_{s} \ge \frac{{\text{S}}}{2}} \\ {0,} & {{\text{otherwise}} } \\ \end{array} } \right.$$
(27)
Alternatively, this can be written using the indicator function as
$$\begin{array}{*{20}c} {\hat{y}_{ens} = \mathop {\arg \max }\limits_{{y \in \left\{ {0,1} \right\}}} \mathop \sum \limits_{s = 1}^{{\text{S}}} {\mathbb{I}}\left( {\hat{y}_{s} = y} \right) } \\ \end{array}$$
(28)
This approach allows healthcare system to reduce computational overhead by parallelizing the training process across nodes while maintaining high prediction accuracy.
link

![Average Cost of Medical School [2025]: Yearly + Total Costs Average Cost of Medical School [2025]: Yearly + Total Costs](https://educationdata.org/wp-content/uploads/2025/09/page-1.png)